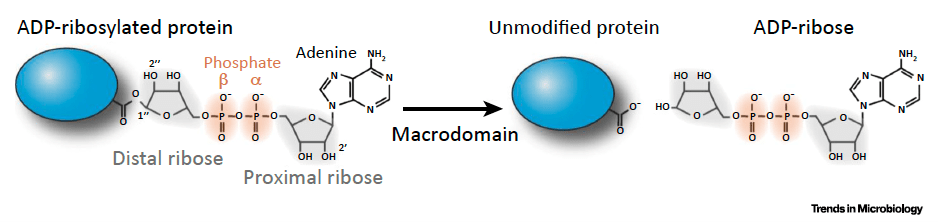
Mono-ADP-ribosylation (MAR) is a common post-translational modification that is generally induced upon virus infection and mediated by cellular enzymes known as PARPs. However, there is still relatively little known about how MAR can inhibit virus replication. Interestingly, coronaviruses (CoVs) all encode for a macrodomain protein (Mac1) which removes MAR from proteins. This indicates that CoVs may be especially sensitive to the antiviral activities of MAR. The Fehr lab is especially interested in the CoV macrodomain, and we have previously discovered that Mac1 counters PARP-mediated MARylation. Using a robust BAC-based reverse genetics platform to make genetic changes to the CoV genome, we have shown that in the absence of Mac1 activity CoVs replicate poorly, especially in vivo, and do not cause disease in animal models of infection. These results demonstrate that the macrodomain is a prominent virulence factor and a potential therapeutic target.

Our primary research interests lie in dissecting the host-virus interactions between cellular PARP enzymes and viral macrodomains. This research includes defining PARPs important for the antiviral response to CoVs, identifying the targets of PARPs and the macrodomain, and determining exactly how the macrodomain promotes virus replication and disease at a mechanistic level. In addition, we are developing novel macrodomain inhibitors that could also be used therapeutically to treat CoV infection. We address these questions using a variety of molecular virology, genetic, and biochemical/chemical biology approaches. Specific projects include:
1. Define the roles and identify mechanisms used by the CoV macrodomain to promote virus replication and block the innate immune response
Our lab uses BAC-based reverse genetic platform to make genetic changes to the CoV genome. This allows us to make specific mutations in viral genes to directly determine the role of that gene in virus replication or other functions during infection. Using this system, we have engineered several mutations in the macrodomain of multiple CoVs (MHV, MERS-CoV, SARS-CoV, SARS-CoV-2) and have determined that the macrodomain promotes virus replication via multiple mechanisms and blocks the production of IFN in certain cells. We have demonstrated that Mac1 uses both its ability to bind to ADP-ribose (reader) and its ADP-ribosylhydrolase (eraser) activity to separately counter PARP-mediated repression of virus replication. We will continue to dissect the mechanisms used by the CoV macrodomain to enhance virus replication and block innate immunity utilizing novel molecular tools to identify Mac1 interacting partners and ADP-ribosylated proteins during infection.
2. Determine the mechanisms PARPs use to inhibit virus infection and promote innate immunity
On the host side, we have developed both PARP knockout mice and cells to aid in determining the role of specific PARPs in CoV infection both in cell culture and in mice. In addition, we utilize siRNA/crRNA knockdowns and specific inhibitors to manipulate the levels and activities of host PARPs during infection. Using these tools, we have demonstrated that i) PARP12 and PARP14 can inhibit CoV infection in the absence of macrodomain activity ii) PARP14 enhances the IFN response following infection and iii) PARP7 restricts IFN production during infection. Our lab is interested in identifying their specific protein targets, determining their role in modulating cellular transcriptional profiles, identifying their mechanisms of action, and identify additional PARPs that repress CoV replication and pathogenesis
3. Understand the biochemistry of the CoV macrodomain

The macrodomain has two distinct biochemical functions, to remove MAR from proteins when attached via acidic residues (hydrolysis), but also to bind to ADP-ribose when it is attached to other residues. Though we have many crystal structures depicting how ADP-ribose fits into the macrodomain via its binding pocket, it is not clear how individual amino acids in this pocket contribute to the overall activity of the macrodomain. We are using recombinant proteins and have developed several different assays to determine how specific residues contribute to either ADP-ribose binding or hydrolysis and then translate that information into how those mutations affect virus replication. So far, we have focused on 4 different residues of Mac1, D22, N40, I131, F132. In general, we’ve found that D22 is critical for ADP-ribose binding, N40 is critical for enzymatic activity, F132 is critical for both activities, and intriguingly I131 seems to control ADP-ribose binding, as mutation of this residue to alanine resulted in enhanced ADP-ribose binding. Surprisingly, enhanced ADP-ribose binding was detrimental for infection. We plan to further dissect the many important residues of this protein to further define how they impact Mac1 biochemistry and virus replication.
4. Discover novel inhibitors of the CoV macrodomain
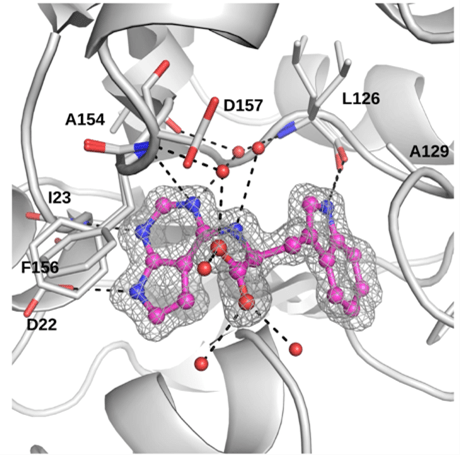
As the CoV macrodomain is clearly essential for these viruses to cause disease, we believe that it may be a therapeutic target that could be important for protecting humans or animals from CoV-related diseases. Using high-throughput assays that measure macrodomain/ADP-ribose binding, we have performed high-throughput screening (HTS) in collaboration with the HTS lab here at KU to identify some compounds that inhibit macrodomain activity in vitro, which we confirmed using multiple assays in collaboration with the Anthony Leung Lab (Johns Hopkins University) and the Lari Lehtio Lab (Oulu University, Finland). Furthermore, worked with Dana Ferraris (McDaniel College) who chemically modified other compounds into potent macrodomain inhibitors. We have demonstrated that several of these Mac1 inhibitors repress CoV replication and we’ve identified drug-resistant mutations that demonstrate the specificity of these compounds and provide unique insight into the biochemistry of Mac1. We continue to use organic synthesis to modify and improve these compounds, develop them into potent anti-viral compounds, and identify additional drug-resistant mutations and determine how these mutations impact the structure and activity of Mac1.
Collaborators
- Rudragouda Channappanavar – Oklahoma State University
- Michael Cohen – Oregon Health Sciences University
- Erik Holmstrom – University of Kansas
- Susan Weiss – University of Pennsylvania
- Lari Lehtiö – Oulu University (Finland)
- Dana Ferraris – McDaniel College (MD)
- Anthony Leung – Johns Hopkins University
- David Davido – University of Kansas
- Robin Orozco – University of Kansas
- Rob Unckless – University of Kansas
- Anuradha Roy – University of Kansas
- Roberto De Guzman – University of Kansas
- Scott Lovell – University of Kansas
- David Johnson – University of Kansas
- Philip Gao – University of Kansas
- Teruna Siahaan – University of Kansas
- Michael Hageman – University of Kansas
- Chris Sullivan – University of Texas
- Masanori Aikawa – Harvard University
- Charles Brenner – City of Hope (CA)
- Adam Hume – Boston University
- Sunil More – Oklahoma State University
- Guy Poirier – Laval University (Canada)
- Ivan Ahel – Oxford University (Great Britain)
- Luis Enjuanes (CNB-CSIC – Spain)
- Stanley Perlman – University of Iowa
